Script to model OPO-Cro-Bgal using Systems Biology Toolbox 2 (http://www.sbtoolbox2.org/main.php)
First draft 9 Dec 2009, Revised 10 Dec, Fixed Cro, W (added dimer diss eq) 11 Dec added intermediate plots for each case to make web publishable
Contents
- OPO Simulations | Input File opo_Bgal091211.txt
- Cro Simulations | Input File opocro091210.txt
- scCro Simulations | Input File opo_scCro091210.txt
- F58W Simulations | Input File opoWcro091210.txt
- Species Plots | PDF summary plot
OPO Simulations
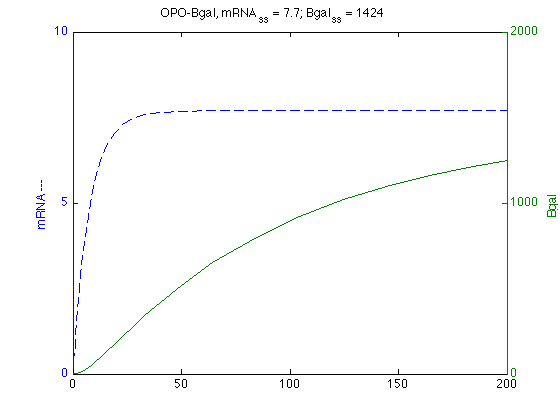
%Read in model from text file. modelO = SBmodel('opo_Bgal091211.txt'); %SBedit(modelO) % allows for interactive editing and simulation, also % The steady-state is given by: % mRNA = 7.69231 % Bgal = 1424.5 % Find max mRNA oSim = SBsimulate(modelO,200); mO=oSim.statevalues(:,1); % FmC=croSim.statevalues(:,1); % Selects 1st column mRNA levels mOmax=max(mO) ; % Returns largest entry % max_mC=find(max(mC)==mC) % Returns index of largest entry mOtmax = oSim.time(find(max(mO)==mO)); % Returns time@max % Plots mRNA on Left and Bgal on Right axis [axo,h1,h2]=plotyy(oSim.time,oSim.statevalues(:,1),oSim.time,oSim.statevalues(:,2)); set(get(axo(1),'Ylabel'),'String','mRNA ---') ; set(get(axo(2),'Ylabel'),'String','Bgal ') ; set(h1,'LineStyle','--') ; % dashed % set(h2,'LineStyle',':') % dotted title ('OPO-Bgal, mRNA_{ss} = 7.7; Bgal_{ss} = 1424')
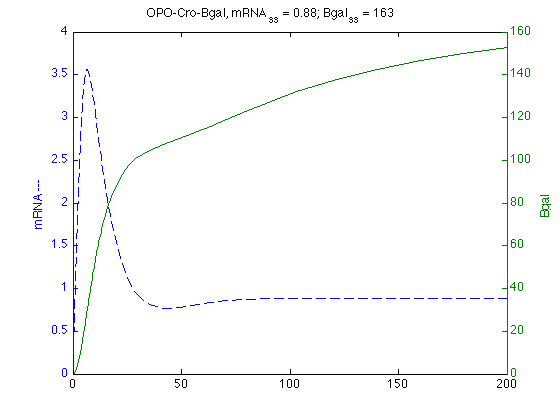
Cro Simulations
%Read in model from text file. modelC = SBmodel('opocro091210.txt'); % SBedit(modelC) % allows for interactive editing and simulation, also % steady state calculations croSim = SBsimulate(modelC,200); % states: {'mRNA' 'Cro' 'Bgal' 'Cro2' 'OPO' 'OPOCro2'} % statevalues: [145x6 double] % SBsteadystate(modelC);% 0.8843 68.2880 163.7647 0.5936 0.1150 0.8850 % Find max mRNA mC=croSim.statevalues(:,1); % Selects 1st column mRNA levels mCmax=max(mC); % Returns largest entry % max_mC=find(max(mC)==mC) % Returns index of largest entry mCtmax = croSim.time(find(max(mC)==mC)); % Returns time@max % Plots mRNA on Left and Bgal on Right axis [axc,hc1,hc2]=plotyy(croSim.time,croSim.statevalues(:,1),croSim.time,croSim.statevalues(:,3)); set(get(axc(1),'Ylabel'),'String','mRNA ---') ; set(get(axc(2),'Ylabel'),'String','Bgal ') ; set(hc1,'LineStyle','--') ; % dashed % set(h2,'LineStyle',':') % dotted title ('OPO-Cro-Bgal, mRNA_{ss} = 0.88; Bgal_{ss} = 163')
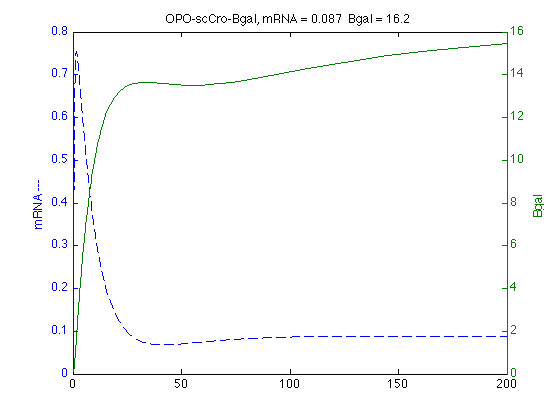
scCro Simulations
modelS = SBmodel('opo_scCro091210.txt'); scSim = SBsimulate(modelS,200); % Find max mRNA mS=scSim.statevalues(:,1); mSmax=max(mS); mStmax = scSim.time(max(mS)==mS); % Plots mRNA on Left and Bgal on Right axis [axs,hs1,hs2]=plotyy(scSim.time,scSim.statevalues(:,1),scSim.time,scSim.statevalues(:,2)); set(get(axs(1),'Ylabel'),'String','mRNA ---') ; set(get(axs(2),'Ylabel'),'String','Bgal ') ; set(hs1,'LineStyle','--') ; % dashed % set(hs2,'LineStyle',':') % dotted title ('OPO-scCro-Bgal, mRNA = 0.087 Bgal = 16.2')
F58W Simulations
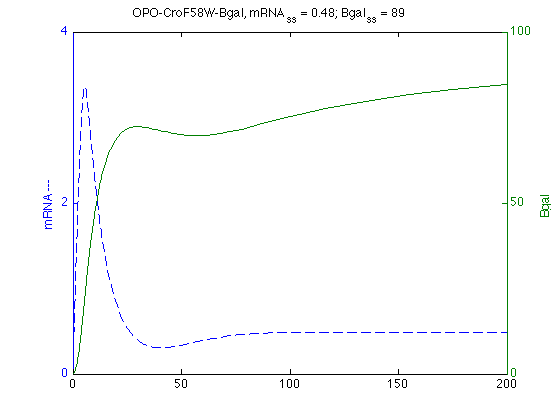
modelW = SBmodel('opoWcro091210.txt'); wSim = SBsimulate(modelW,200); % Find max mRNA mW=wSim.statevalues(:,1); mWmax=max(mW); mWtmax = wSim.time(max(mW)==mW); % Plots mRNA on Left and Bgal on Right axis [axw,hw1,hw2]=plotyy(wSim.time,wSim.statevalues(:,1),wSim.time,wSim.statevalues(:,3)); set(get(axw(1),'Ylabel'),'String','mRNA ---') ; set(get(axw(2),'Ylabel'),'String','Bgal ') ; set(hw1,'LineStyle','--') ; % dashed % set(hw2,'LineStyle',':') % dotted title ('OPO-CroF58W-Bgal, mRNA_{ss} = 0.48; Bgal_{ss} = 89')
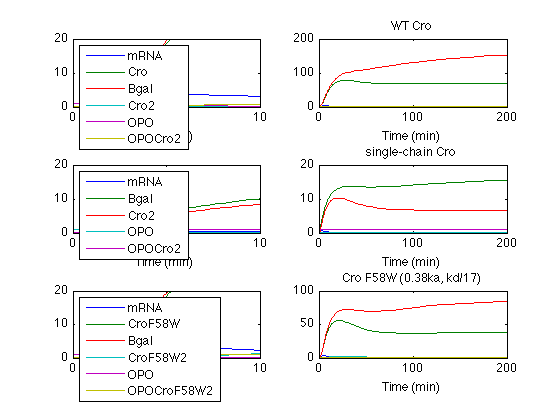
Species Plots
wt Cro short 50 = 3.6
subplot(3,2,1) % hcs = plot (croSim.time(1:50), croSim.statevalues(1:50, :)) hcs = plot (croSim.time, croSim.statevalues); axis([0 10 0 20]) legend (croSim.states,'Location','Northwest') xlabel('Time (min)') % wt Cro full subplot(3,2,2) hc = plot (croSim.time, croSim.statevalues); title('WT Cro') xlabel('Time (min)') % legend (croSim.states,'Location','EastOutside') % scCro short 100= 4 units subplot(3,2,3) % hss = plot (scSim.time(1:100), scSim.statevalues(1:100, :)) hss = plot (scSim.time, scSim.statevalues); axis([0 10 0 20]) xlabel('Time (min)') legend (scSim.states,'Location','Northwest') % scCro subplot(3,2,4) hs = plot (scSim.time, scSim.statevalues); title('single-chain Cro') xlabel('Time (min)') % legend (scSim.states,'Location','EastOutside') % Cro F58W short 50= 4.3 subplot(3,2,5) % hws = plot (wSim.time(1:50), wSim.statevalues(1:50, :)) hws = plot (wSim.time, wSim.statevalues); axis([0 10 0 20]) xlabel('Time (min)') legend (wSim.states,'Location','Northwest') % Cro F58W subplot(3,2,6) hw = plot (wSim.time, wSim.statevalues); title('Cro F58W (0.38ka, kd/17)') xlabel('Time (min)') % legend (wSim.states,'Location','EastOutside') set(gcf, 'PaperPosition', [0.5 0.5 8 10.5]); print -dpdf sims091211.pdf